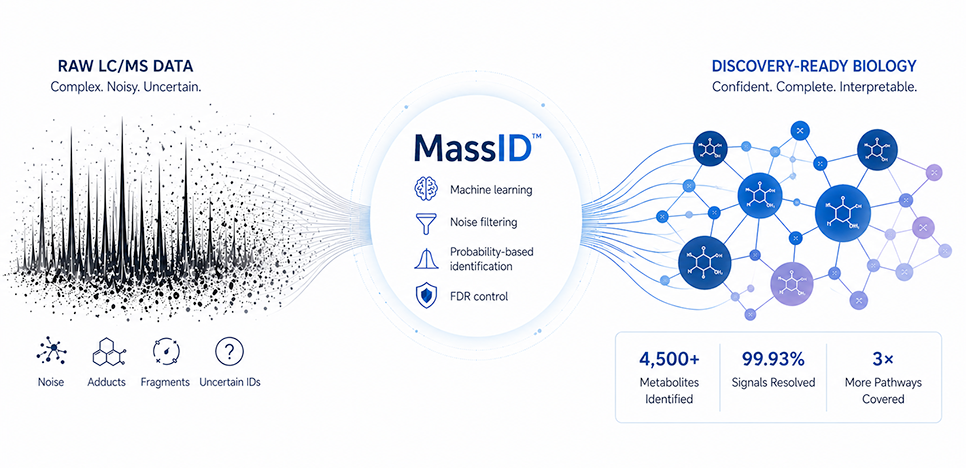
What if untargeted metabolomics delivered on its promise?
For years, untargeted metabolomics has been one of the most compelling approaches in biology: measure broadly, capture molecular diversity and improve biological interpretation and understanding. In practice, however, it often results in noisy data, uncertain identifications and outputs that are difficult to interpret with confidence.
Panome Bio has addressed this challenge by improving analytical rigor in untargeted workflows. With MassID™, that framework has been further extended.
The Promise (and Problem) of Untargeted Metabolomics
Metabolites are closely linked to phenotype, reflecting the combined effects of genetics, diet, environment and lifestyle. This makes metabolomics a powerful tool for studying biological systems. Modern LC/MS technologies can detect thousands of molecular signals in a single run. However, most analytical workflows still rely on targeted strategies that:
- Focus on a few hundred known metabolites
- Exclude the majority of detected signals
- Apply strict filtering to reduce false positives
While these approaches increase confidence, they substantially limit biological coverage.
Untargeted Data: Rich but Complex
Panome Bio’s Next-Generation Metabolomics® captured all detectable signals, including unknown compounds. But raw LC/MS datasets are inherently complex and include:
- Redundant signals (adducts, fragments)
- Contaminants and technical artifacts
- Ambiguous or overlapping identifications
Without robust computational methods, researchers face a tradeoff between retaining data depth and ensuring reliability.
The MSI Problem: Confidence Without Quantification
The Metabolomics Standards Initiative (MSI) introduced qualitative levels of identification confidence:
- Level 1: confirmed with reference standards
- Level 2: high-quality spectral match
- Level 3: compound class assignment
While widely used, MSI levels have important limitations. Even Level 1 identifications can be incorrect, particularly for isomers or structurally similar compounds where mass, fragmentation and retention time are not fully discriminative. More importantly, MSI does not provide a quantitative estimate of identification accuracy. There is no associated probability or false discovery rate. As a result, analyses often rely on conservative filtering, reducing dataset size and limiting biological interpretation.
Enter MassID: Confidence and Coverage
MassID was built to remove that tradeoff entirely. Instead of asking researchers to choose between depth and reliability, it delivers both, by fundamentally changing how metabolite identification works. Resulting in cleaner, more reliable data.
At its core, MassID combines:
- Machine learning–driven signal detection
- Advanced noise and artifact filtering
- robability-based identification models
- Explicit false discovery rate (FDR) control
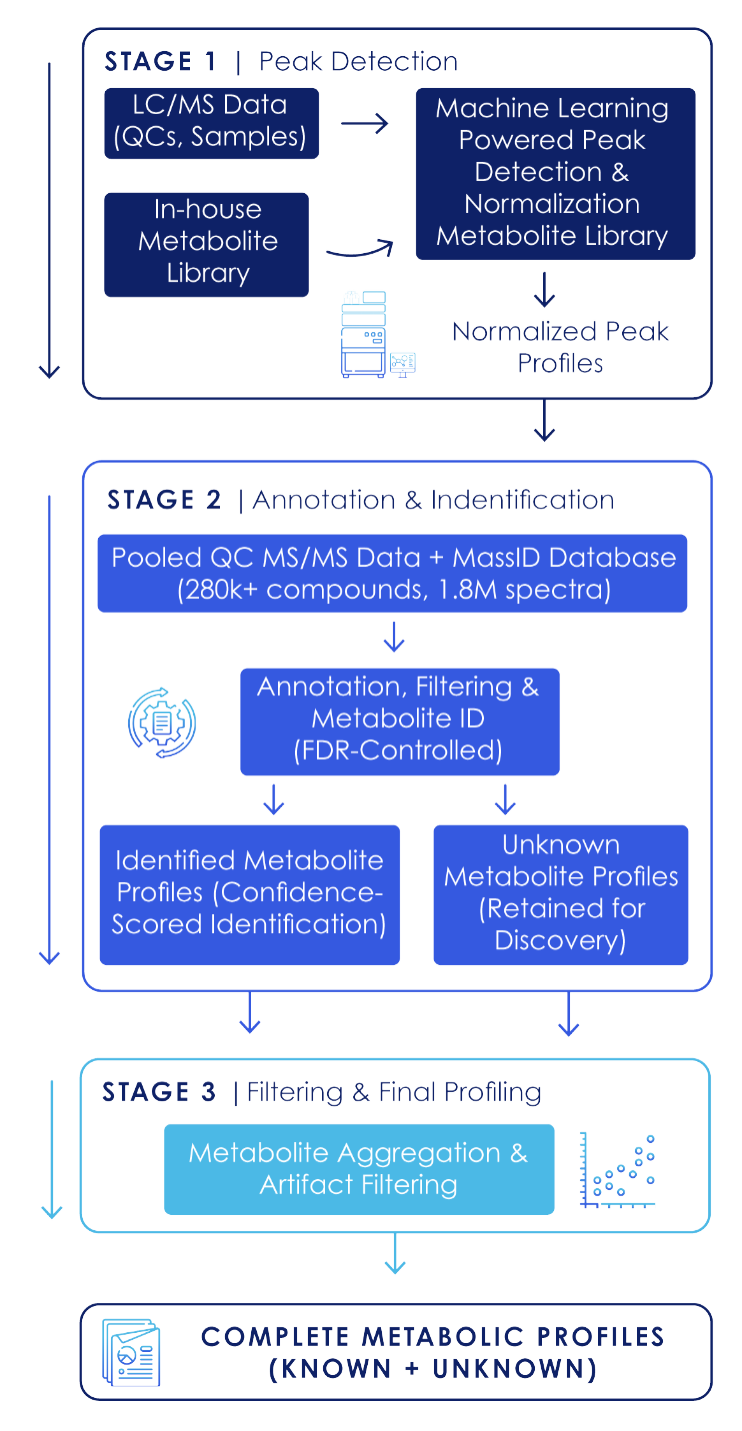
Figure 1: MassID Workflow. MassID performs machine learning based signal detection, normalization, noise removal, and probability-based identification with FDR control to generate complete, discovery-ready metabolic profiles.
From Raw Data to Interpretable Profiles
MassID applies a structured workflow to transform LC/MS data into analysis-ready profiles:
- Signals are detected with high sensitivity using machine learning
- Data is aligned, normalized, and corrected across runs
- Noise, contaminants and redundant features are removed
- Unstable signals (high variability) are filtered out
- Remaining features are assigned quantitative confidence scores
Importantly, even unknown metabolites are retained and incorporated into the overall metabolic profile, preserving the full structure of the metabolome.
A Quantitative Approach to Confidence
Instead of labeling metabolites with vague categories, MassID assigns a probability of being correct.
This allows researchers to:
- Apply objective thresholds (e.g. 5% FDR)
- Keep more data while maintaining statistical rigor
- Evaluate uncertainty directly
And for tricky cases like isomers? MassID groups indistinguishable structures and combines their probabilities, ensuring chemically meaningful results for downstream analysis.
Built on a Comprehensive Reference Framework
Accurate identification depends on both computational methods and high-quality reference data. MassID is powered by a massive and continuously growing database:
- 280,000+ known compounds
- 1.8 million fragmentation spectra
- Integrated public and proprietary resources
In addition, experimentally measured retention times from more than 1,000 compounds are used to train predictive models. These models estimate chromatographic behavior across different separation methods, providing an additional layer of evidence during identification.
Application to Human Plasma
When applied to human plasma from patients with cardiovascular disease (CVD), a complex biological matrix, MassID demonstrated the following:
- nearly 200,000 detected features
- 4,500+ structurally identified metabolites
- Systematic removal of noise and artifacts
- 99.93% of signals classified as either identified metabolites or noise
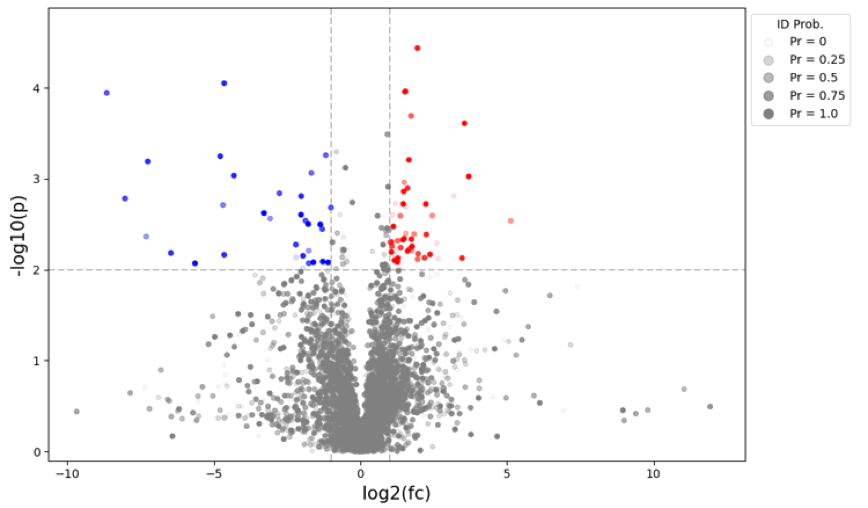
Figure 2: Application of MassID to Human Plasma in CVD. Volcano plot highlights differential metabolites by probability. Statistical analysis shows log2(fc) and -log10(p) values. Each dot is a metabolite. Dots are shaded according to the identification probability.
For context: Traditional approaches typically identify fewer than 800 metabolites. Even under stringent filtering (5% FDR), MassID still retains substantially more metabolite features than MSI-based workflows.
Expanded Biological Coverage
Including all confidently identified metabolites leads to broader biological interpretation:
- Pathway coverage increased from 63 to 193 pathways
- That’s more than a threefold expansion
This enables a more complete view of metabolic networks that would be missed using conventional filtering approaches.
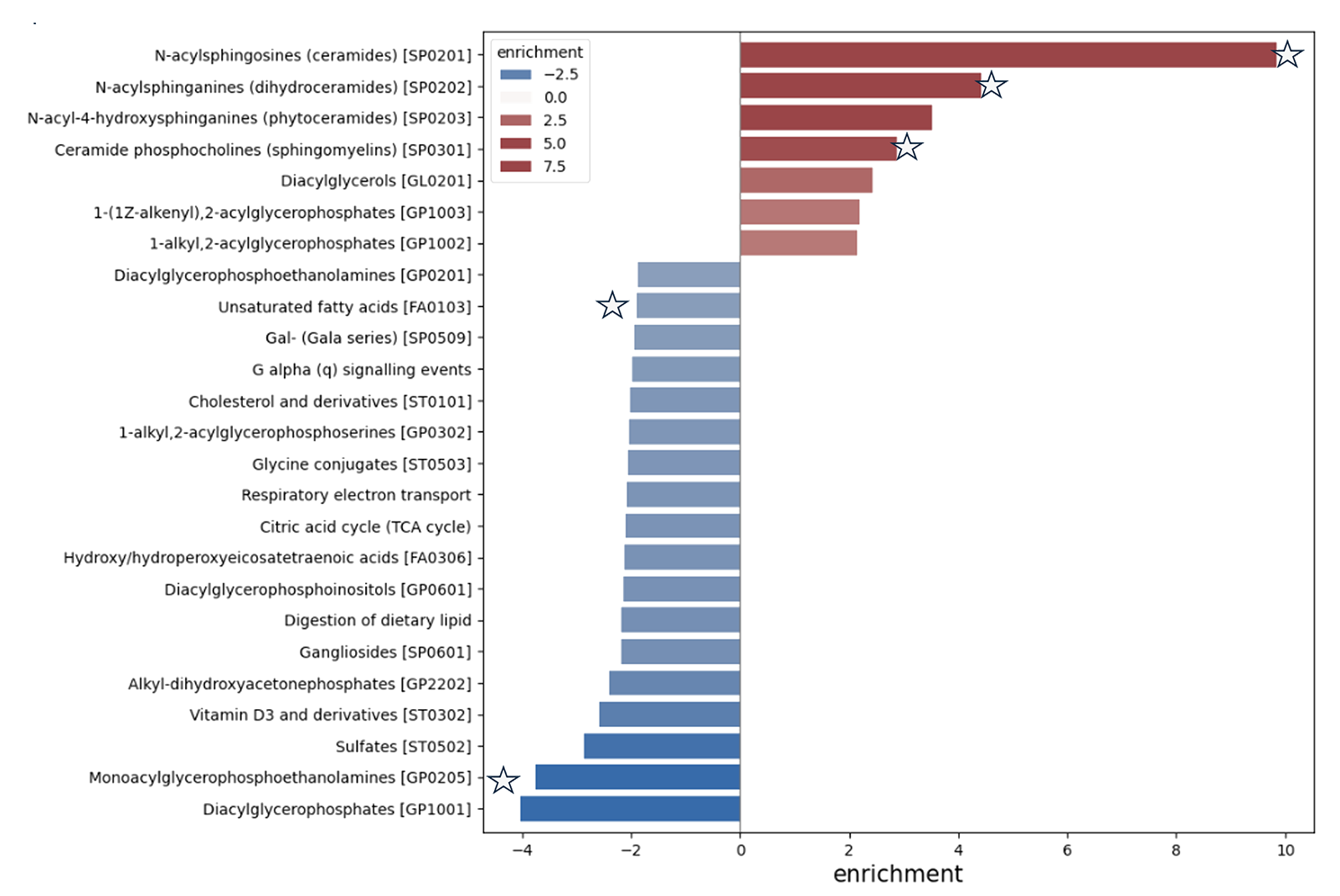
Figure 3: Pathway enrichment analysis with MassID in CVD. Pathway enrichment analysis on the right, red indicates upregulated and blue downregulated metabolites in CVD.
The Bottom Line
Untargeted metabolomics has long been limited by challenges in identification confidence and data interpretation. MassID addresses these challenges through probability-based identification, FDR control and improved data curation. This allows researchers to work with datasets that are both comprehensive and statistically well-defined, without sacrificing biological coverage.